Post Analysis Tutorial¶
Outline¶
This quick tutorial will guide you through the creation of an additional gene-set on a pre-existing Enrichment Map. It can for example help to localize microRNA, transcription factors or drug targets in enriched pathways displayed on the map. The new gene-set that we want to add to the network is called the signature gene-set.
Download the test data: PostAnalysisTutorial.zip
Description of the tutorial files contained in the PostAnalysisTutorial folder:
CTCF_DIFF.gmtSignature gene-set.Human_GO_AllPathways_no_GO_iea_April_15_2013_symbol.gmtGene-set file used to create the original Enrichment MapES12_EM_example.cysThe Enrichment Map on which we want to add the signature gene-set
Instructions¶
- Open Cytoscape and Open ES12_EM_example.cys
- Click on Plugins / Enrichment Map/ Post Analysis
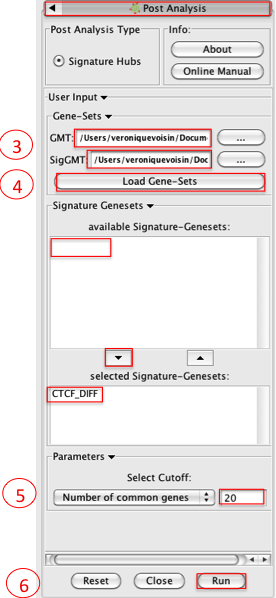
- Please select the following files by clicking on the respective (…) button and selecting the file in the Dialog:
- GMT / ‘Human_GO_AllPathways_no_GO _iea_April_15_2013_symbol.gmt’
- SigGMT / ‘CTCF_DIFF.gmt’
- Click on Load Gene-Sets
- In the Signature-Genesets box: click on CTCF_DIFF as available Signature-Genesets
- Click on the down arrow to move CTCF_DIFF in the lower box
- Tune parameters
- Set Number of common genes to 20
- Click on Run
Examining Results¶
ES12_EM_example_PA_CTCF_Diff.cys
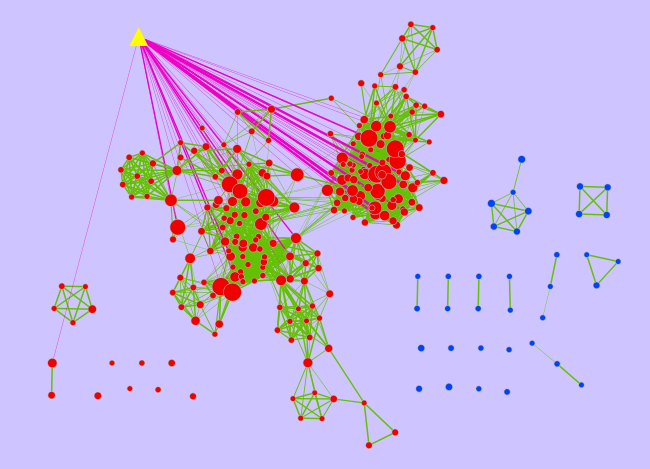
- The yellow triangle is the signature gene-set and the pink edges represent overlap of 20 genes or more between the signature gene-set and a given gene-set from the Enrichment Map (red node). The width of the pink edges is proportional to the number of genes in the overlap.
- This signature gene-set represents the CTCF (transcription factor) target genes that are different in MCF7 cells treated or not with Estrogen.
- Clicking on a pink edge will show the genes contained in this overlap in the gene expression panel:
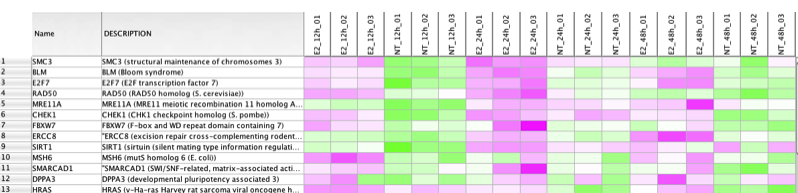
Note
The gene-signature gene-set corresponds to ENCODE CHIP-seq data for the cell line MCF-7 treated or not with estrogen and for the CTCF factor using the tool CSCAN/Browse data (http://159.149.160.51/cscan/). The signature gene-set includes the genes that were different between the two conditions (treated or not with estrogen)